LEB on GitHub
Downloads
Event-driven acquisition (EDA)
Event-driven acquisition is a framework for adaptive microscopy integration in Micro-Manager. It allows for neural network-based image analysis to be used to detect events of interest in real-time and control microscope acquisition parameters accordingly.
SPARTAN
SPARTAN is a MATLAB-based software package specialised on particle-type datasets. While SPARTAN allows to perform general SMLM analysis steps (localization, drift correction and channel registration), it can perform particle averaging or simulation tasks and provides an interface with established EM 3D reconstruction routines. Cite (4).
B-Store
B-Store is a lightweight format for storing SMLM data and metadata in a HDF5 container. In addition, it provides a number of Python analysis routines for efficiently and quickly processing high-throughput SMLM datasets.
FIFI SimMLA
Fourier optics simulation code for laser-excitation microscopes utilizing dual microlens arrays. This is handy if you want to set up FIFI on your high NA-objective microscope. See https://github.com/kmdouglass/simmla. Cite (3).
AutoLase
MicroManager plugin for real time closed loop control of the density of photoactived molecules during a PALM measurement. Enables automated PALM imaging. Should come packaged with MicroManager. See https://micro-manager.org/wiki/AutoLase and (1).
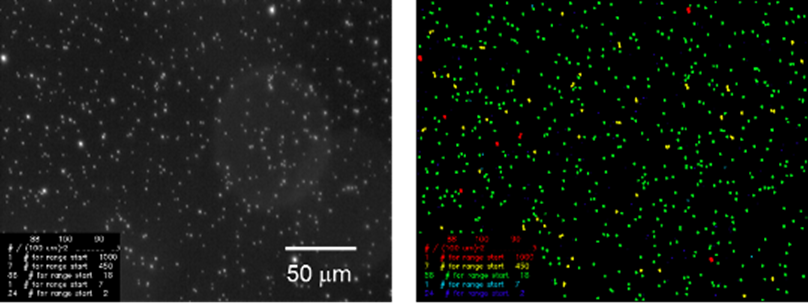
High throughput PALM
MicroManager plugin for automated acquisition of many fields of view (FOVs) during a PALM experiment. Focused on microbiological samples, it searches for FOVs containing sufficient numbers of bacteria in the phase contrast channel, and then takes a PALM measurement. It can use AutoLase to control the density of photoactivated molecules during the PALM measurement. Cite (1).
PALMsiever
MATLAB software for the visualization and analysis of single-molecule localization microscopy data. It includes interactive browsing, filtering and analysis, plus the ability to extend its functionality through plugins. Cite (2).
Other resources

Please cite the relevant publication if you use our software in work leading to a publication.
1. SJ Holden et al. (2014) High throughput 3D super-resolution microscopy reveals Caulobacter crescentus in vivo Z-ring organization. Proc Natl Acad Sci 201313368.
2. T Pengo et al. (2015) PALMsiever: a tool to turn raw data into results for single-molecule localization microscopy. Bioinformatics 31(5):797-798.
3. K Douglass et al. (2016) Super-resolution imaging of multiple cells by optimized flat-field epi-illumination. Nat. Photonics 10.1038.
4. C Sieben*, N Banterle*, et al. (2018) Multicolor single-particle reconstruction of protein complexes. Nat Methods 10.1038/s41592-018-0140-x.